MITIE is the Modular Inter-Layer Feedback Topology InvEstigation Tool and Simulator by me, Jason Spencer.
The original version was used to run many of the simulations used for my PhD thesis "An Investigation into the Scale-free Nature of Heterogeneous Networks", published 2008. The current version is a major re-write as the original was created for "back of the laptop" simulations and prototyping. This version is heavily re-factored with extensibility and speed in mind, but with the tried, matured and tested algorithms of the original.
This page is arranged into the following sections:
Additionally please see Core library details for information about the core libraries and How to extend the functionality of MITIE for information about how to extend MITIE and create new modules.
MITIE is a program that allows the user to specify node and link evolution strategies for a network and to examine the properties of the network during the evolution.
It is a simulation tool that performs a series of epoch-driven operations that either alter or analyse a network topology. These operations, or processes as they are referred to in MITIE, can consider flows and available network resources, and thus can be used to model the effect of inter-layer feedback. MITIE is built on an easily extendable framework which could be used to add very specific functionality that isn't already available.
MITIE was created to simulate multi-layer telecommunications networks but could be used to simulate many kinds of transport networks, including road transport networks (highways, vehicle level, container level), biological (the cardio-vascular system), ecological networks and so on.. The network design algorithms available in MITIE are mostly stochastic in nature, but there are also deterministic algorithms available in the MITIE framework.
To create a 1000 node topology from an initial chain of 10 nodes (that is ten vertices joined in a row by nine bi-directional edges) and then grow the network by adding a node and two links into the existing network, linking them with a preference for nodes of a higher degree:
jsp@nostromo:~/mitie$ ./mitie -initChain 10 -maxSize 1000 '0:grow{D,const=1}' '0:linkNodes{prev={add,new},D,const=1}' 'END:weightsDump{label=finalweight}' > doc/sampleoutput1.txt
jsp@nostromo:~/mitie$ cat doc/sampleoutput1.txt | grep "^finalweight"|cut -sf 2- - > doc/sampleoutput1.adj
The output of MITIE is deliberately verbose and designed for further parsing. The full output can be seen in file doc/sampleoutput1.txt . The label argument to the weightsDump process option specifies a label to be prepended to each line of weightsDump output. This can be grepped, and the prefix removed, to create a final weights matrix - as in doc/sampleoutput1.adj
Just for sanity, checking the degree distribution of the adjacency matrix (this could be done with the degDistDump process in MITIE, which outputs a degree distribution instead of a weights matrix) to check for an approximate power-law..
jsp@nostromo:~/mitie$ cat doc/sampleoutput1.adj | gawk '{ node_degree=0; for(i=1;i<=NF;++i) if($i!=0) ++node_degree; print node_degree; }' | sort -n | uniq -c | tee doc/sampleoutput1.dd.dat
482 2
208 3
103 4
61 5
40 6
22 7
16 8
10 9
8 10
<output snipped>
And then graphed and fitted to a power-law by gnuplot..
jsp@nostromo:~/mitie$ gnuplot -p -e "FIT_LIMIT = 1e-24; f(x) = a * x ** b; fit f(x) 'doc/sampleoutput1.dd.dat' using 2:1 via a,b; set log xy; set xlabel 'outdegree'; set ylabel 'frequency'; plot f(x) title sprintf(\"%0.3f x ** %0.3f\",a,b), 'doc/sampleoutput1.dd.dat' using 2:1 title 'MITIE output'"
Which looks like ..
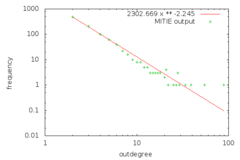
[enlarge to SVG] | [enlarge to PNG]
To examine the effect of network erosion on the top 10 eigenvalues of an input topology, where the erosion is based on removing the least routed over links:
jsp@nostromo:~/mitie$ ./mitie -initWt doc/sampleoutput1.adj '0:remLink{rpl}' '0:eigenvalDump{comb_lap,desc,count=10,label=eigval_,append_epoch}' > doc/sampleoutput1.eig.txt
jsp@nostromo:~/mitie$ cat doc/sampleoutput1.eig.txt | grep "^eigval_" | head -n 5
eigval_0 90.0742 56.1002 40.1699 35.1942 31.0387 29.6085 29.2999 28.1421 27.706 25.9598
eigval_1 90.0742 56.1002 40.1691 35.1942 31.0387 29.6085 29.2999 28.1421 27.706 25.9598
eigval_2 90.0741 55.0959 40.1691 35.1941 31.0386 29.6062 29.2781 28.1421 27.706 25.9597
eigval_3 90.0739 55.0959 40.1691 35.1941 30.9799 29.5639 29.276 27.9891 27.0129 25.9596
eigval_4 90.0738 55.0959 40.1691 35.1941 30.9799 29.5639 29.276 27.9891 27.0129 25.9596
The input file, doc/sampleoutput1.adj, is the grepped output from Example 1 which is this time loaded as an initial weights matrix, and the processes being performed at every epoch are remLink and eigenvalDump. The rpl (Routes Per Link) option selects a link, preferrentially selecting the least routed over links (that is the links with the least number of shortest paths traversing them (irrespective of traffic pattern)). The eigenvalDump options specify that the eigenvalues of the combinatorial Laplacian adjacency matrix are to be calculated, and count specifies how many of them are to be output. "desc" specifies that they are to be sorted in descending numerical order before being output.
The output is sent to doc/sampleoutput1.eig.txt.
Then, to plot the effect of link erosion on the largest eigenvalue of the topology:
jsp@nostromo:~/mitie$ cat doc/sampleoutput1.eig.txt | grep "^eigval_" > doc/sampleoutput1.eig.dat
jsp@nostromo:~/mitie$ cat doc/sampleoutput1.eig.txt | grep "^eigval_" | gawk -F[_\ ] '{print 1989-$2 " " $3 }' | gnuplot -p -e "set term png size 1200, 800; set output \"figure2.png\"; set xrange [2000:1000]; set yrange [0:100]; set xtics 100; set xlabel 'Number of links'; set ylabel 'Largest Eigenvalue'; plot '-' title 'Link erosion' "
The 1989 figure is a conversion factor from the epoch number which is in the MITIE output to the number of remaining links. The graph then looks like..
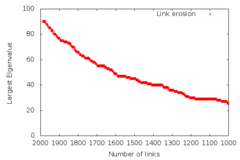
[enlarge to SVG] | [enlarge to PNG]
Because of the command-line nature of MITIE all of the above can be easily run multiple times (and distributed over multiple hosts) to get useful error bounds.
In general MITIE is implemented in such a way that it does the work on the network, and the analysis is done by other software which is better suited to it - under UNIX awk, perl, sed etc.. are your friends, under Windows MATLAB may be a good option, or cygwin.
The project is arranged into two parts - the main executable and the core libraries that contain the processes, link/node selectors, which perform the useful actions on the topology layers.
mitie, mitie_static or mitie.exe is the main executable - mitie (the version that uses DLLs to store the core libraries) and mitie_static (a statically linked version) for Linux, and mitie.exe for Windows. The differences between the versions for each platform are described in the section Windows vs. Linux versions , below. mitie looks in ./libs for the .so files to load.
The main executable performs the following tasks, in order:
This is a collection of modules that perform operations on the topology. A module can be one of the following types:
Network Process This type of module directly performs operations on a topology. See Processes for more details. Examples of network processes include: node selector, and the link is assigned a weight generated by the specified dimensioner link selector and assigning a weight to the link which was generated with a dimensioner Link Selector This type of module selects, depending on the contect, either an existing link in the topology, or a pair of nodes which aren't already connected and may be connected (with a network process such as addLink). See Link Selectors for more details. Examples include: Node Selector This type of module selects a node from the topology for use in the operation of a Network Process. See Node Selectors for more details. Examples include: Link Dimensioner (or just Dimensioner) is a module that generates a new weight for a link. The link could be an existing link, in which case a dimensioner could be invoked for the purposes of re-dimensioning, or a new link. See Link Dimensioners for more details. Examples include:
A full list of modules in the core libraries is available on the page Core library details.
MITIE can be built with the GNU C++ (ver 4.4.5 tested under Ubuntu Linux and an unknown version tested under CentOS), LLVM/CLang (ver. 2.8 under Linux tested)(unfortunately g++ is still required for automatic dependency generation in make) or Microsoft Visual Studio (2008 and 2010 tested). Additionally, the following libraries are either required or optional:
Boost is required - specifically Boost.Filesystem, Boost.mpl, Boost.Random, Boost.Test, Boost.lexical_cast, Boost.shared_ptr, Boost.logic, Boost.algorithm.string and Boost.ScopeExit
GSL - The GNU Scientific Library is optional - used only for eigenvalDump at the moment.
Doxygen is optional - used to create these docs, so it's only needed if you've updated the docs.
GraphViz/DOT is optional - used by doxygen to make prettier class diagrams.
The command line syntax is the same for Windows and Linux versions of MITIE. Start MITIE with --help for a more detailed description of the available options, but a typical command line may look like:
./mitie -initChain 10 -maxSize 5000 '0:grow{D,const=1}' 'END:distDistDump{}'
This creates an empty network and then intialises it to a chain of ten nodes (that is ten nodes are connected by nine (bidirectional) links in a line/chain, i.e. node 0 is connected to node 1, which is also connected to node 2 etc..). A maximum size is specified for the network which is 5000 nodes. The following processes are instantiated and added to the process list:
'0:grow{D,const=1}'
This specifies that beginning with epoch 0, and repeated at every epoch until infinity the grow process is to be executed. There are two arguments to the process, D happens to be a node selector which will select the node to connect the new added node to, and const=1 which invokes the const link dimensioner with an argument of 1, which tells grow to assign a link weight of 1 to the link connecting the newly added node.
Because grow increases the network size the process above is repeated until no new nodes can be added - this happens when the network size reaches the specified maximum (5000 nodes from -maxSize). Once this occurs, any processes specified with a timing of END will be invoked.
'END:distDistDump{}'
This process is invoked at the end of the simulation - in this case the process called distDistDump finds the shortest paths through the topology and reports the frequency of the sum of weights on the minimum cost path.
The general syntax describing a process on the command line is as follows:
start[+lifetime][@step][,prio]:proc_name{[option1][,option2..]}
or
END[,prio]:proc_name{[option1][,option2..]}
Where:
start is the first epoch in which to perform the process. If this is "END" then the process is performed at the end of the simulation. lifetime (optional) if the number of epochs for which to repeat the process. If not specified this is assumed to be infinite. step (optional) The rate at which to repeat the process. If not specified then it is assumed to be 1, i.e. repeat every epoch until the lifetime expires. prio (optional) The priority of the process. Processes with a lower value priority will be performed before processes with a higher value. Processes with the same priority will be performed in the same order in which they appear in the command line. If not specified the priority is assumed to be 0. proc_name The name of the process to perform. option1,option2.. A list of comma separated options which will be passed to the process.0:hop{} 0:skip{} 0:jump{}
Would perform the processes hop, skip, jump repetitatively, in that order, from the first epoch until a termination condition occurs.
0+11,2:jump{} 0+10:hop{} 0+10:skip{} END:relax{}
Would perform the processes hop, skip, jump in that order (although jump appears first, it has a priority of 2), for the first ten epochs, and then a jump in the eleventh, and then end the epoch cycling, and call relax.
-initChain 10 -maxSize 100 '0:grow{ D, const=1 }' '0@2:linkNodes{ prev=new, D, const=1 }' 'END:adjLDump{}'
As the command line options to mitie would initialise the topology to a chain of ten nodes, and then at every epoch add a new node and connect it to an existing node (with a link of weight 1 - see const), selecting it at random but preferring high degree nodes (see D); on the first, third, fifth etc.. epoch (epoch number 0,2,4,...) it would also add a second link between the newly added node (see prev) and a second existing node, again preferring higher degree nodes, and again with link weight 1. Once the network grows to 100 nodes the epoch loop will terminate and the adjacency list will be printed (see adjLDump).
It is important to use single quotes (') on Unix shells to escape the process configuration string otherwise the curly brackets {} will be expanded by the shell. Under Windows the quotes are also recommended to keep the process description as a single command line argument.
The versions are mostly identical apart from the following differences
MITIE has a single Makefile which has a number of targets. make the usage target to get a full list of available targets:
jsp@fatman:~/Desktop/XP share/mitie$ make usage Available build targets.. Debug builds generate much more verbose debugging info, may contain additional sanity checks, disable compiler optimisations and include GDB info mitie - build a dynamically linked version of the main mitie executable (remember to build corelibs or corelibs_dbg also) mitie_dbg - build a dynamically linked debug version of the main mitie executable (remember to build corelibs_dbg or corelibs also) mitie_static - build a statically linked version of the main mitie executable mitie_dbg_static - build a statically linked debug version of the main mitie executable corelibs - builds the core DLLs corelibs_dbg - builds the core DLLs as a debug build libs/mitie_core_np.so - builds mitie_core_np.so only libs/mitie_core_np_dbg.so - builds mitie_core_np_dbg.so only libs/mitie_dump_np.so - builds mitie_dump_np.so only libs/mitie_dump_np_dbg.so - builds mitie_dump_np_dbg.so only libs/mitie_core_dim.so - builds mitie_core_dim.so only libs/mitie_core_dim_dbg.so - builds mitie_core_dim_dbg.so only libs/mitie_core_nsel.so - builds mitie_core_nsel.so only libs/mitie_core_nsel_dbg.so - builds mitie_core_nsel_dbg.so only libs/mitie_core_lsel.so - builds mitie_core_lsel.so only libs/mitie_core_nsel_dbg.so - builds mitie_core_nsel_dbg.so only run_unit_tests - run all of the unit tests Unit test targets: CjMatrix_test - Basic test for CjMatrix CjMatrix_unit_test - Unit test for CjMatrix CjNLTopology_unit_test - Unit test for CjNLTopology CjNetworkProcess_unit_test - Unit test for CjNetworkProcess CjNetworkProcessFactory_unit_test - Unit test for CjNetworkProcessFactory CjNetworkProcessFactory_unit_test_static - Unit test for CjNetworkProcessFactory, but with the core libs statically compiled in unit_tests - build all unit tests Miscellaneous targets: all - build all libs, tests and mitie doc - Run Doxygen to generate html source code documentation tar - Build tar file of source and testing files clean - delete temporary build files, .so output files, all variants of the mitie binary (debug, static etc..) jsp@fatman:~/Desktop/XP share/mitie$
Invoking make clean and then make mitie corelibs is a typical way of building. (make -j 2 mitie corelibs is recommended on a dual core machine) CXX in the Makefile can be switched from g++ to clang and should work fine.
The Visual C++ under Windows build process has many less options - a simple nmake /f mitie.mak is all that can be done.
Since MITIE only supports static linking at the moment, there are no other options for building and no need to build corelibs separately. It may be necessary to update the location of the Boost libraries in the INCOPTS variable in mitie.mak
The command line options can be found by specifying --help in the command line:
jsp@nostromo:~/mitie$ ./mitie --help
System is Linux nostromo 2.6.35-30-generic #61-Ubuntu SMP Tue Oct 11 15:29:15 UTC 2011 i686
Start time is Thu Nov 10 21:26:28 2011
Got 1 CLI options: --help
./mitie --help [process, dimensioner, link or node selector name]
./mitie --help_all
./mitie --desc_procs
./mitie [options] [process description]
where [options] can be one or more of:
-initWt <file> Specifies a file containing the initial link weights
-loadGeo <file> Specifies a file containing the geography of the nodes.
-maxSize N Specifies the maximum size of the network (N nodes).
-initChain N Initialises the initial topology to a chain of N nodes.
-initMesh N Initialises the initial topology to a fully connected mesh of N nodes.
-seed SEED Specifies an integer, SEED, with which to seed the random number generator, otherwise the seed is derived from the current time.
-maxCycles N Specifies the maximum number of epochs/cycles to perform in the simulation.
Each process can be described as:
start[+lifetime][@step][,prio]:proc_name{[option1][,option2..]}
END[,prio]:proc_name{[option1][,option2..]}
Where:
start is the first epoch in which to perform the process. If this is "END" then the process is performed at the end of the simulation.
lifetime if the number of epochs for which to repeat the process. If not specified this is assumed to be infinite.
step The rate at which to repeat the process. If not specified then it is assumed to be 1, i.e. repeat every epoch until the lifetime expires.
prio The priority of the process. Processes with a lower value priority will be performed before processes with a higher value. Processes with the same priority will be performed in the same order in which they appear in the commend line. If not specified the priority is assumed to be 0.
proc_name The name of the process to perform.
option1,option2.. A list of comma separated options which will be passed to the process.
Network processes registered: { adjLDump degDistDump distDistDump distancesDump eigenvalDump pajekDump routesDump topStatDump weightDistDump weightsDump } { addLink remLink } grow linkNodes redim
Node selectors registered: D EP EW R RPN node prev
Link selectors registered: degree geoabs geopow geowax random rpl wt
Link dimensioners registered: const nroutes
End time is Thu Nov 10 21:26:28 2011
jsp@nostromo:~/mitie$
To get list of library modules with a brief one-line description call MITIE with the --desc_procs switch (typical output). For detailed information on a module call MITIE with the --help <module_name> switch. To obtain detailed information for all modules call MITIE with the --help_all switch (typical output).
MITIE does not do much file handling since it is designed to be used in a command line context with external text handling tools for analysis. The input files that MITIE does read, however, are through the -initWt and -loadGeo switches. An input file contains a list of numbers delimited by space, horizontal tabs or a comma "," and by carriage returns (CR or CR/LF). Lines beginning with a #(hash,pound,octothorp) are considered comments and ignored.
-initWt requires a square matrix of integers equal to or greater than 0. i.e. the number of space/tab/comma delimited integers per line is the same as the number of non-comment lines.
-loadGeo requires either:
Output file formats are dependant on the source - at the moment this is only the "dump" network processes - these can forward their output to a file instead of STDOUT so the format is the same as may be found from Common output options. If you would like to store the state of the final weights matrix to re-use in a new invocation of MITIE then use a similar grep and cut as in Example 1.
There is plenty I'd like to do with MITIE, and I welcome feature requests. My current best thinking on what to implement next is:
Version 1.0 07 Jun 2009 - The initial release
Version 1.1 23 Oct 2009 - Added eigenvalDump and pajekDump. Added lazy bridge calculation - some operations like repetitative link removal (network erosion) are now up to 36% faster.
 1.7.1
1.7.1